You can help by funding the sequencing and analysis of 3,000 genomes.
$100 Donation or more:
Goes toward analysis of a gene sequence.
$250 Donation or more:
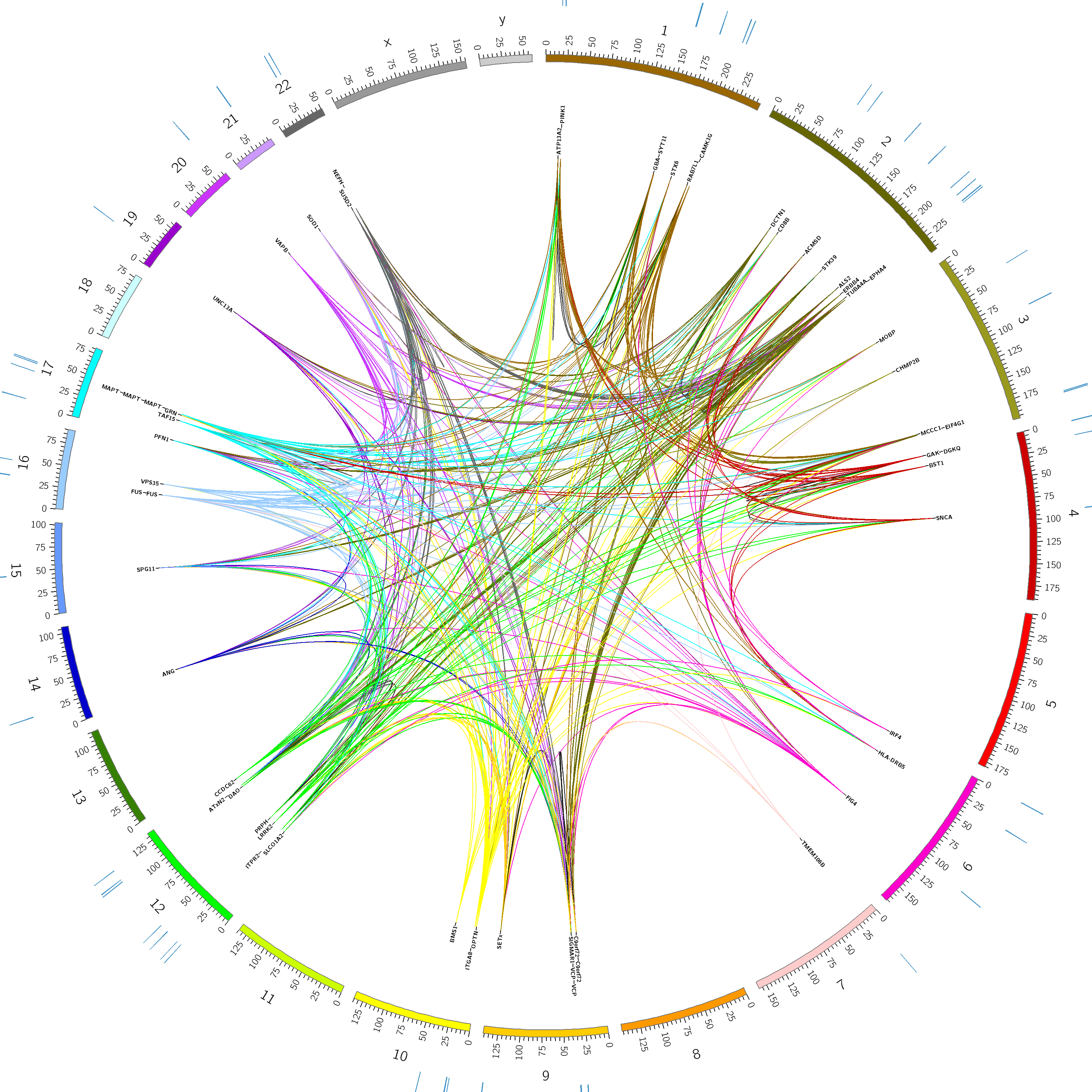
Receive a unique and beautiful framed genome plot of one of our sequenced genes.
$1,500 Donation:
Pays for one full gene sequencing. Receive a set of five different framed genome plots.
Why the time is ripe for a push on research into the genetics of PSP
- 4 years ago we were paying over $1,000 to sequence the “exome” – the portion of the genome containing the genes that code proteins — representing about 3% of our DNA.
- Today we can sequence the whole genome — and identify disease-related changes that may reside between genes — at a similar cost. We can generate ~30-fold more data and provide a comprehensive look at an individual patient’s entire genome.
- Currently, there are around 2,000 samples available for sequencing. New samples are being accrued at the rate of 150-200 per year.
- This resource is unique. It is unlikely that another group would be able to create a similar resource of validated PSP cases for genetic analysis.
- Once high quality whole genome sequences are completed, the data collection phase for those samples will be finished—i.e., there will not be ‘missing data’ or a better methodology requiring re-analysis in the future. The DNA sequence, once determined, can be used in multiple analyses and tested against other samples in an ever increasing database—but the data we generate today will be a core and unchanging foundation for all future sequence analysis and annotation studies of PSP.
- The targeted sample size—at least 2,000 whole genomes, and potentially as many as 3,000 whole genomes from PSP patients will provide adequate statistical power to identify new risk alleles and rare, (putative) disease-causing mutations. It is unlikely that any future collection of samples will match or exceed the statistical power of the collection in the current consortium.
- For instance, the ENCODE project (Encyclopedia of DNA Elements) has suggested that 75-80% of DNA regions have a biologic function—with the minimum functional attribute being transcription into RNA. ENCODE has generated a wealth of information on gene regulation and functional elements residing between genes that will inform analysis of the PSP whole genome sequences.
- Reference databases such as 1000 genomes, DBGAP and others are either available for comparative analysis or will come online during the data analysis phase of the PSP genetics project.
- Gerald Schellenberg and colleagues at Penn
- Dan Geschwind and colleagues at UCLA
- Owen Ross and colleagues at Mayo Clinic, Florida
- John Hardy and colleagues of University College, London UK
- Gunther Hoeglinger of DZNE, Munich, Germany
These investigators have run large-scale sequencing and analysis projects in related disorders such as Alzheimer’s disease, Huntington’s Disease, Parkinson’s Disease and Autism. Their groups maintain expertise and infrastructure required to support sequence generation, sequence analysis pipelines, database creation and targeted analysis of large scale sequence databases.
- Data sharing, including access to database for non-consortium investigators who propose novel analyses
- Joint analysis of data with publications based on new findings to include all consortium members
- Commitment to promote the careers of junior investigators who make significant contributions to the consortium
- The PSP Genetics Consortium draws on the scientific, financial and managerial strengths of both CurePSP and the Tau Consortium
- This allows for a project of broader scope than either organization could pursue alone
- Additional resources will allow for more samples to be sequenced and analyzed more rapidly—accelerating the pace of discovery of novel genetic targets that will:
- Provide additional information on genetic changes that increase risk of developing PSP
- Identify additional targets for development of therapeutics to treat PSP
- Provide leads for the development of new diagnostic approaches to PSP